Paper companion notebook — Jones et al.
Auto-generated from
tutorial_notebook/notebook_paper.ipynb. Executed against the seaborn canvas so every figure is inline as a static PNG. Plotly-only cells are kept for context and marked as placeholders — for the interactive version, run the source notebook.
ethoscopy & ethoscope-lab: a framework for behavioural analysis to lower entrance barrier and aid reproducibility
Figure 1
1. Loading the data
import ethoscopy as etho
import pandas as pd
# This tutorial required version 1.1.8 or greater
etho.__version__
'2.0.5'
One-time setup: fetch the tutorial datasets
The tutorial pickle files (~36 MB total, dominated by overview_data.pkl) are not bundled with the PyPI package, to keep pip install ethoscopy lean.
Run the cell below once per environment to download them into the installed package directory. Subsequent runs are idempotent (already-present files are skipped).
You can also fetch them manually from https://github.com/gilestrolab/ethoscopy/tree/main/src/ethoscopy/misc/tutorial_data.
# Idempotent: skips files that are already present.
import ethoscopy as etho
etho.download_tutorial_data()
[skip] overview_data.pkl (already present)
[skip] overview_meta.pkl (already present)
[skip] circadian_data.pkl (already present)
[skip] circadian_meta.pkl (already present)
[skip] 4_states_F_WT.pkl (already present)
[skip] 4_states_M_WT.pkl (already present)
Tutorial data ready in: /home/gg/.cache/ethoscopy/tutorial_data
PosixPath('/home/gg/.cache/ethoscopy/tutorial_data')
# import this function to get the tutorial dataset
from ethoscopy.misc.get_tutorials import get_tutorial
# We'll be using the same dataset as in the overview tutorial, so use the same function below with the argument 'overview'
# Load the data and metadata, and then intialise it into a behavpy_HMM
data, metadata = get_tutorial('overview')
df = etho.behavpy(data, metadata, check = True)
df = df.t_filter(start_time = 24, end_time = 144)
(a)
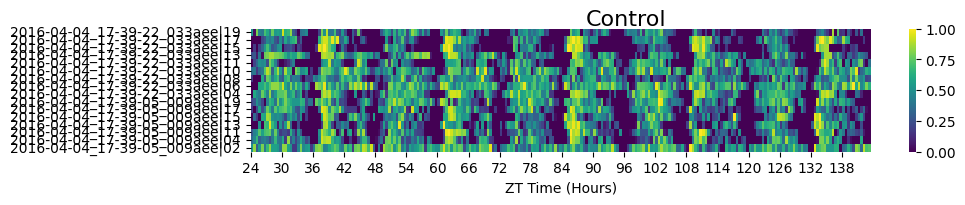
fig = df.xmv('sleep_deprived', False).heatmap(variable = 'asleep', title = 'Control')
fig.show()
/home/gg/Code/ethoscope_project/ethoscopy/src/ethoscopy/behavpy_core.py:1611: FutureWarning: DataFrameGroupBy.apply operated on the grouping columns. This behavior is deprecated, and in a future version of pandas the grouping columns will be excluded from the operation. Either pass `include_groups=False` to exclude the groupings or explicitly select the grouping columns after groupby to silence this warning.
tdf.groupby("id", group_keys=False).apply(
/tmp/ipykernel_557712/1229634965.py:2: UserWarning: FigureCanvasAgg is non-interactive, and thus cannot be shown
fig.show()
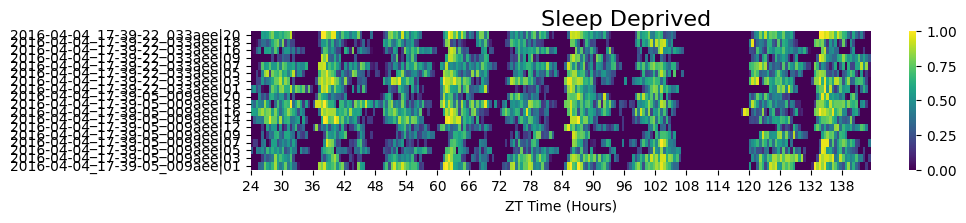
# As we saw in the overview tutorial the dataset we have has half the specimens sleep deprived later in the experiment
fig = df.xmv('sleep_deprived', True).heatmap(variable = 'asleep', title = 'Sleep Deprived')
fig.show()
/home/gg/Code/ethoscope_project/ethoscopy/src/ethoscopy/behavpy_core.py:1611: FutureWarning: DataFrameGroupBy.apply operated on the grouping columns. This behavior is deprecated, and in a future version of pandas the grouping columns will be excluded from the operation. Either pass `include_groups=False` to exclude the groupings or explicitly select the grouping columns after groupby to silence this warning.
tdf.groupby("id", group_keys=False).apply(
/tmp/ipykernel_557712/762731294.py:3: UserWarning: FigureCanvasAgg is non-interactive, and thus cannot be shown
fig.show()
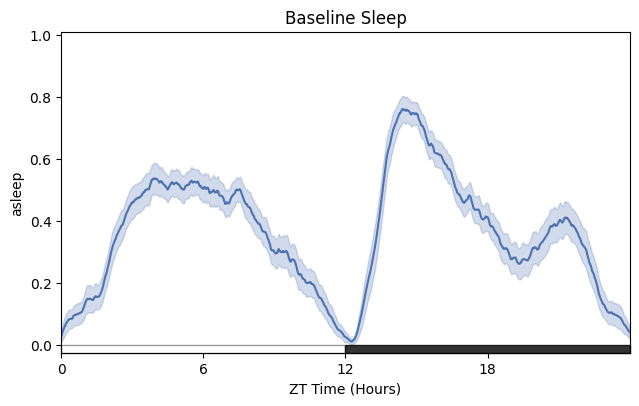
(b)
fig = df.t_filter(start_time = 24, end_time = 96).plot_overtime(variable = 'asleep', wrapped = True, title = 'Baseline Sleep')
fig.show()
/tmp/ipykernel_557712/211641461.py:2: UserWarning: FigureCanvasAgg is non-interactive, and thus cannot be shown
fig.show()
(c)
# Ethoscopy also has pretrained HMMs for both Male and Female CS Drosophila, trained on several days worth of data on hundreds of flies
from ethoscopy.misc.get_HMM import get_HMM
# Have the argument as "M" or "F" for the male or female dataset respectively
h = get_HMM("M")
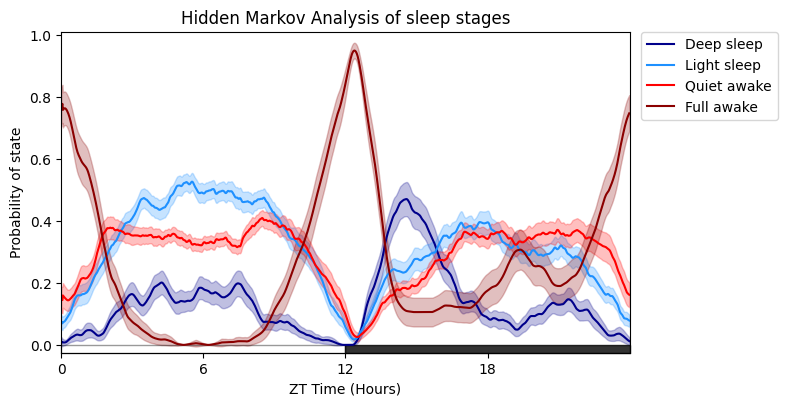
fig = df.plot_hmm_overtime(
hmm = h,
variable = 'moving',
wrapped = True,
labels = ['Deep sleep', 'Light sleep', 'Quiet awake', 'Full awake'],
t_bin = 60,
title = 'Hidden Markov Analysis of sleep stages'
)
fig.show()
/home/gg/Code/ethoscope_project/ethoscopy/src/ethoscopy/behavpy_core.py:1611: FutureWarning: DataFrameGroupBy.apply operated on the grouping columns. This behavior is deprecated, and in a future version of pandas the grouping columns will be excluded from the operation. Either pass `include_groups=False` to exclude the groupings or explicitly select the grouping columns after groupby to silence this warning.
tdf.groupby("id", group_keys=False).apply(
/tmp/ipykernel_557712/1927658757.py:9: UserWarning: FigureCanvasAgg is non-interactive, and thus cannot be shown
fig.show()